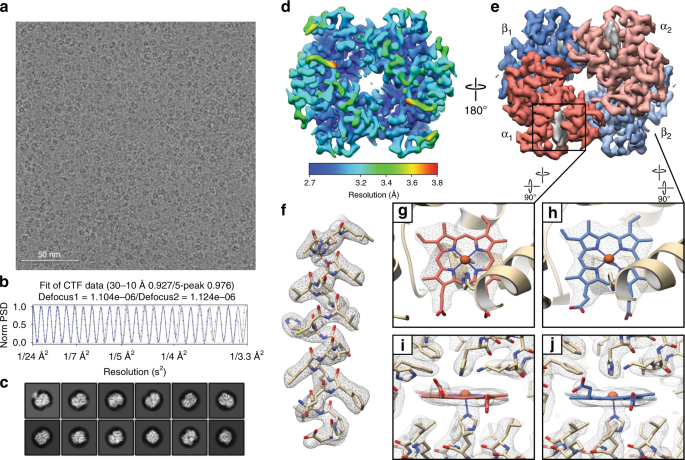
High-resolution structure determination of sub-100 kDa complexes using conventional cryo-EM | Nature Communications
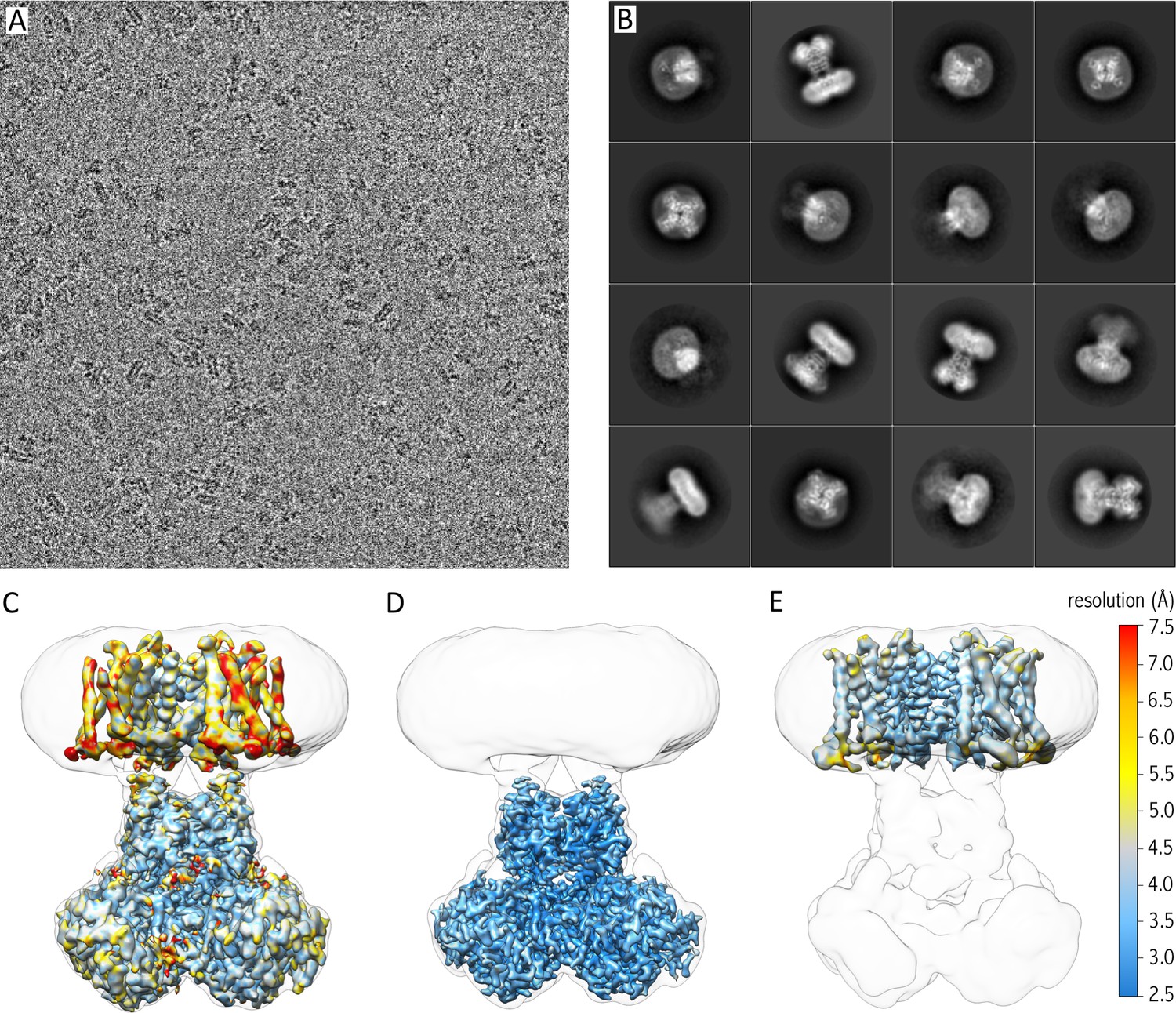
Single-particle cryo-EM structure of a voltage-activated potassium channel in lipid nanodiscs | eLife

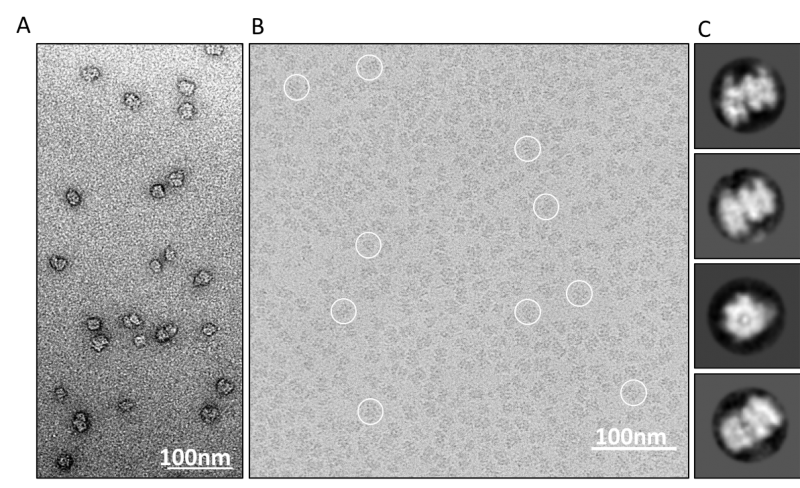
Protein structure by optimized negative staining (OpNS) EM. Electron... | Download Scientific Diagram

Cryo-EM structure of the tetracycline resistance protein TetM in complex with a translating ribosome at 3.9-Å resolution | PNAS
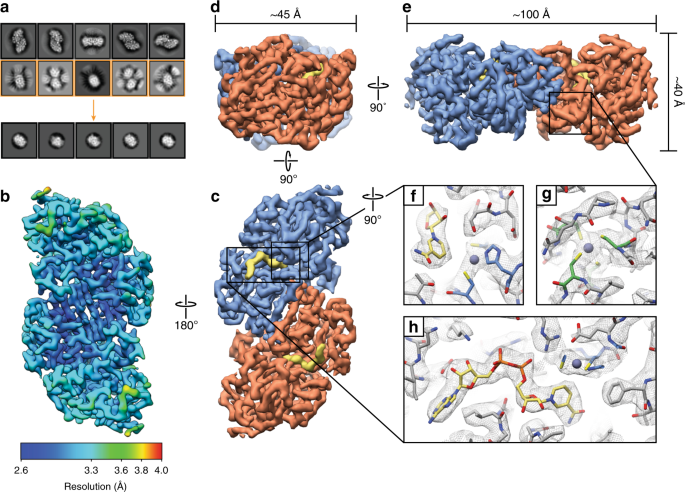
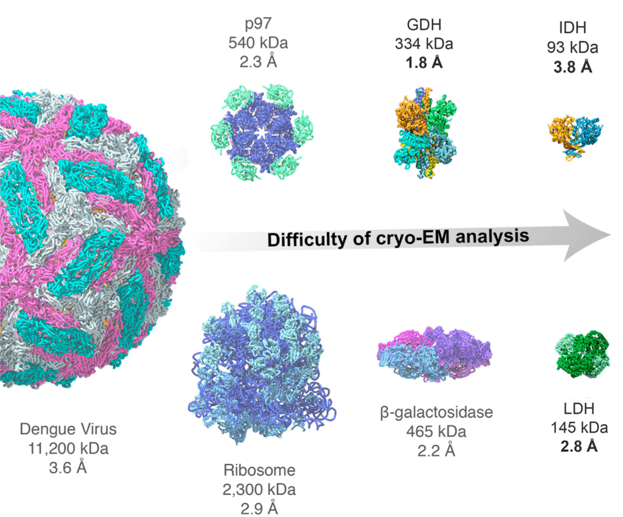
High-resolution structure determination of sub-100 kDa complexes using conventional cryo-EM | Nature Communications

Cryo-EM structures of two proteins in SMALPs. (A) AcrB-SMALP from E.... | Download Scientific Diagram
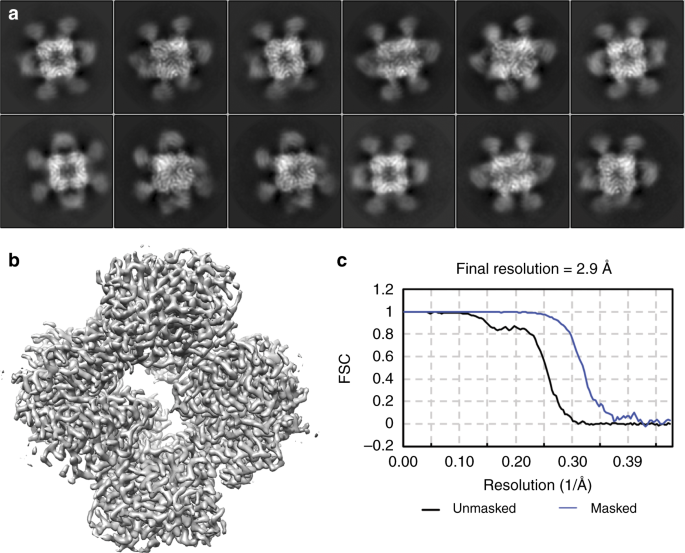
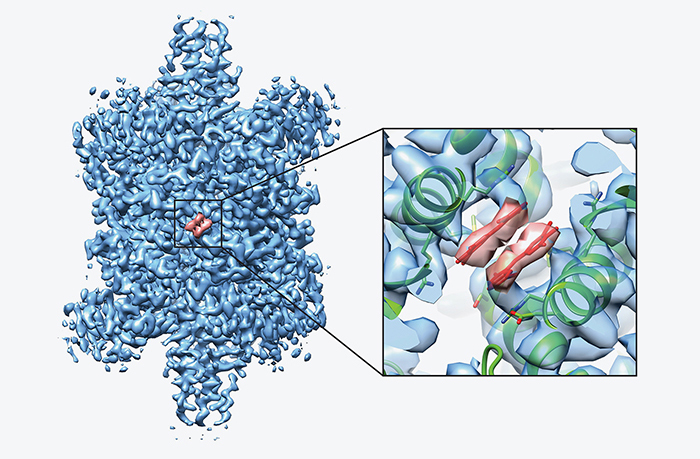
A 3.8 Å resolution cryo-EM structure of a small protein bound to an imaging scaffold | Nature Communications